Tidy summarizes information about the components of a model. A model component might be a single term in a regression, a single hypothesis, a cluster, or a class. Exactly what tidy considers to be a model component varies across models but is usually self-evident. If a model has several distinct types of components, you will need to specify which components to return.
Arguments
- x
An
pamobject returned fromcluster::pam()- col.names
Column names in the input data frame. Defaults to the names of the variables in x.
- ...
Additional arguments. Not used. Needed to match generic signature only. Cautionary note: Misspelled arguments will be absorbed in
..., where they will be ignored. If the misspelled argument has a default value, the default value will be used. For example, if you passconf.lvel = 0.9, all computation will proceed usingconf.level = 0.95. Two exceptions here are:
See also
Other pam tidiers:
augment.pam(),
glance.pam()
Value
A tibble::tibble() with columns:
- size
Size of each cluster.
- max.diss
Maximal dissimilarity between the observations in the cluster and that cluster's medoid.
- avg.diss
Average dissimilarity between the observations in the cluster and that cluster's medoid.
- diameter
Diameter of the cluster.
- separation
Separation of the cluster.
- avg.width
Average silhouette width of the cluster.
- cluster
A factor describing the cluster from 1:k.
Examples
# load libraries for models and data
library(dplyr)
library(ggplot2)
library(cluster)
library(modeldata)
data(hpc_data)
x <- hpc_data[, 2:5]
p <- pam(x, k = 4)
# summarize model fit with tidiers + visualization
tidy(p)
#> # A tibble: 4 × 11
#> size max.diss avg.diss diameter separation avg.width cluster
#> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <fct>
#> 1 3544 13865. 576. 15128. 93.6 0.711 1
#> 2 412 3835. 1111. 5704. 93.2 0.398 2
#> 3 236 3882. 1317. 5852. 93.2 0.516 3
#> 4 139 42999. 5582. 46451. 151. 0.0843 4
#> # ℹ 4 more variables: compounds <dbl>, input_fields <dbl>,
#> # iterations <dbl>, num_pending <dbl>
glance(p)
#> # A tibble: 1 × 1
#> avg.silhouette.width
#> <dbl>
#> 1 0.650
augment(p, x)
#> # A tibble: 4,331 × 5
#> compounds input_fields iterations num_pending .cluster
#> <dbl> <dbl> <dbl> <dbl> <fct>
#> 1 997 137 20 0 1
#> 2 97 103 20 0 1
#> 3 101 75 10 0 1
#> 4 93 76 20 0 1
#> 5 100 82 20 0 1
#> 6 100 82 20 0 1
#> 7 105 88 20 0 1
#> 8 98 95 20 0 1
#> 9 101 91 20 0 1
#> 10 95 92 20 0 1
#> # ℹ 4,321 more rows
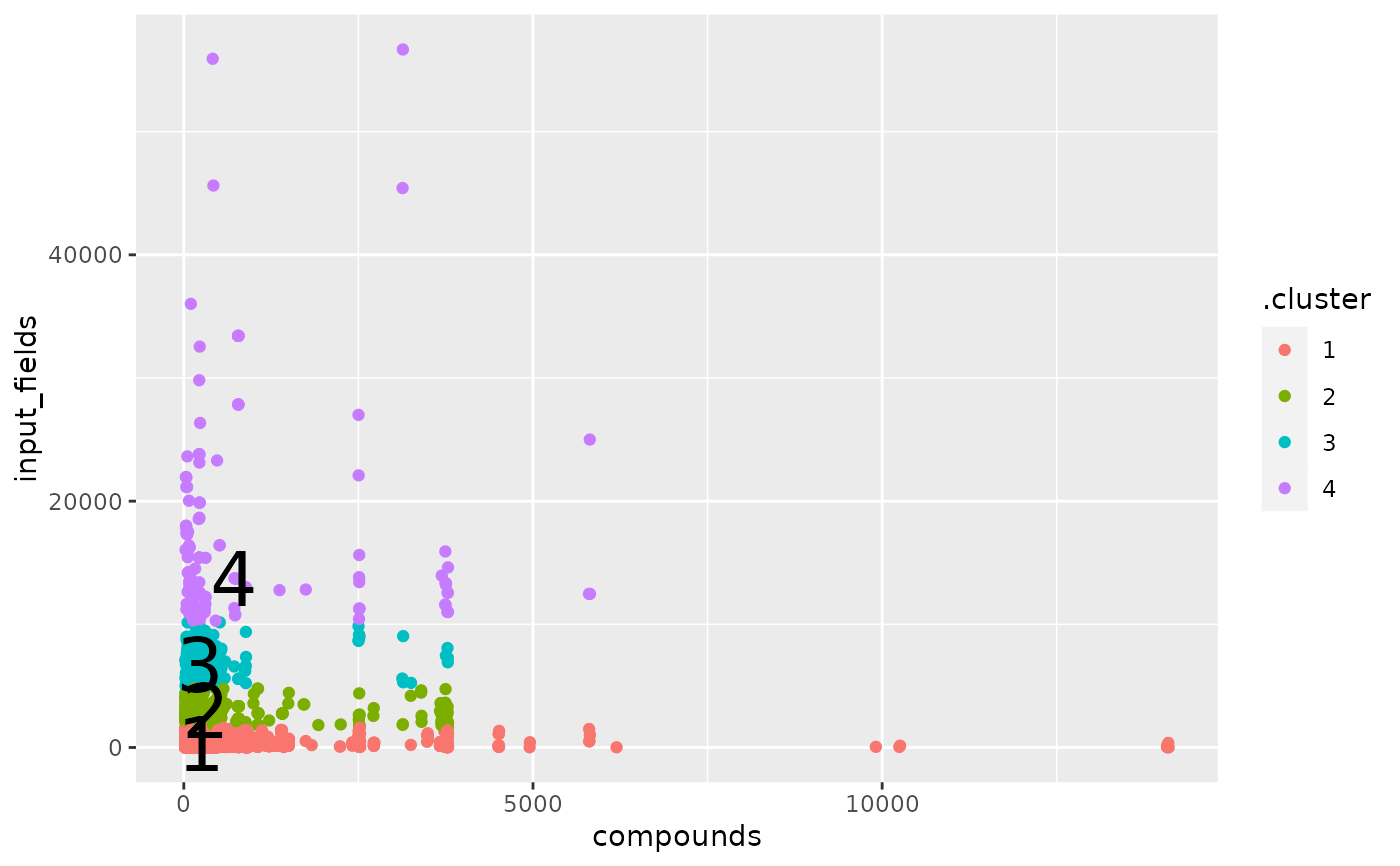
augment(p, x) |>
ggplot(aes(compounds, input_fields)) +
geom_point(aes(color = .cluster)) +
geom_text(aes(label = cluster), data = tidy(p), size = 10)